Intermediate Python Tutorials
Ready to go beyond the basics? These tutorials teach you the skills you need to write more structured, professional Python code. If you’re comfortable with variables, loops, functions, and basic data structures, you’re in the right place.
If you’re new to Python, start with our Python Basics tutorials first.
Note: Browse all resources below, or commit to a structured sequence of guided intermediate Learning Paths with progress tracking.
When you’re ready to go deeper, check out our advanced Python tutorials.
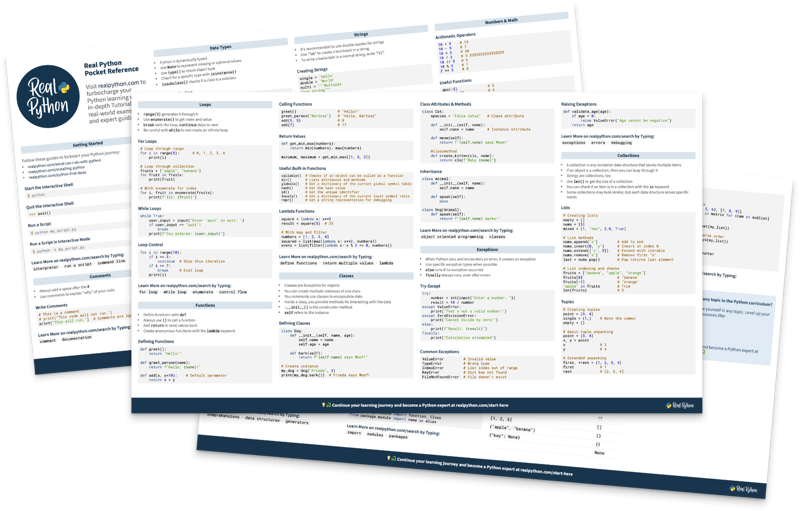
Free Bonus: Python Cheat Sheet
Get a Python Cheat Sheet (PDF) and learn the basics of Python 3, like working with data types, dictionaries, lists, and Python functions:

Not Sure Where You Stand?
Take the Python Skill Test to find out which level fits you best:
Take the Quiz: Test your knowledge with our interactive “Python Skill Test” quiz. You’ll receive a score upon completion to help you track your learning progress:
Interactive Quiz
Python Skill TestTest your Python knowledge in a skills quiz with basic to advanced questions. Are you a Novice, Intermediate, Proficient, or Expert?
You should be comfortable with Python fundamentals like variables, data types, loops, conditionals, functions, and basic data structures such as lists and dictionaries. If you can write simple scripts and understand how to import modules, you’re ready for intermediate content. If any of that sounds unfamiliar, start with our basics tutorials first.
Basic Python covers the core building blocks of the language. Intermediate Python takes you deeper into topics like object-oriented programming, error handling, file I/O, working with APIs, virtual environments, testing, and writing more structured code. You’ll move from writing scripts to building real applications.
Yes! Follow our intermediate learning paths for a structured sequence with progress tracking. If you prefer to explore on your own, start with object-oriented programming and work your way into topics like decorators, generators, and testing.
With intermediate skills, you can build web applications, command-line tools, data pipelines, API integrations, and automation scripts. Check out our project tutorials for hands-on ideas that put your new skills to work.
When you’re comfortable with classes, decorators, context managers, error handling, and testing, you’re ready for the next step. If you can design and structure a multi-file Python project on your own, head over to the advanced tutorials.